How to Read Control/not-digested Plasmids After Electrophoretic Separation
Congratulations, you take a plasmid expressing your gene of involvement (YGOI) and are set to swoop into your functional experiments! Whether you've cloned the plasmid yourself or obtained it from a colleague down the hall, it is always a good thought to take some fourth dimension to confirm that you are working with the correct construct, and verify that the plasmid you received matches the expected sequence. Here at Addgene, we use NGS-based quality control to ostend the sequence of all the plasmids we distribute. This method is time-intensive, so we recommend a variety of means to screen and verify your plasmids. Here, we'll cover brake digest assay. Diagnostic digests tin can exist used to confirm the rough structure of the plasmid based on the predicted sizes and arrangement of different featuresinside the plasmid. Brake analysis can also be used successfully even if you don't accept the full plasmid sequence. One time you have purified plasmid DNA, this method can be done right in your lab in less than a day. Diagnostic restriction digests are comprised of 2 separate steps: one) incubating your Dna with restriction enzymes which cleave the DNA molecules at specific sites and two) running the reaction on an agarose gel to determine the relative sizes of the resulting Deoxyribonucleic acid fragments. Restriction digests are ordinarily used to confirm the presence of an insert in a particular vector by excising the insert from the backbone. To do this, you'll use enzymes with brake sites that flank the insert. You volition need to know both the approximate size of the vector courage as well every bit the predicted size of the insert. You can search NCBI for YGOI to find the particular reference sequence if necessary. Watch this video for a quick overview of how to analyze a brake assimilate: The following tips will make it easier for y'all to obtain a useful and informative diagnostic restriction digest. More Plasmid Eductional Resources: 
Diagnostic restriction assimilate
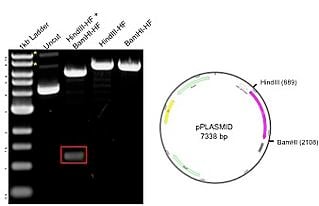 The case plasmid on the correct has a total size of vii.3kb, including a 1.2 kb insert. The plasmid was digested with two unique enzymes (HindIII and BamHI) and run on an agarose gel. The resulting gel image includes a 1kb ladder (lane one) that has bands ranging from about 500bp to 10kb, with the 3.0kb fragment having increased intensity to serve equally a reference band. The uncut Dna (lane 2) shows 3 possible plasmid conformations, with relaxed and nicked marked with asterisks (*). When the plasmid is digested with eitherHindIII and BamHI solitary (lanes four-5), there is a single band of 7.three kb representing the full size of the plasmid. The double digest with both HindIII and BamHI (lane 3) produces bands at 6kb and 1.2kb (red box), matching the backbone and insert, respectively. The results on the gel correspond to the predicted sizes.
The case plasmid on the correct has a total size of vii.3kb, including a 1.2 kb insert. The plasmid was digested with two unique enzymes (HindIII and BamHI) and run on an agarose gel. The resulting gel image includes a 1kb ladder (lane one) that has bands ranging from about 500bp to 10kb, with the 3.0kb fragment having increased intensity to serve equally a reference band. The uncut Dna (lane 2) shows 3 possible plasmid conformations, with relaxed and nicked marked with asterisks (*). When the plasmid is digested with eitherHindIII and BamHI solitary (lanes four-5), there is a single band of 7.three kb representing the full size of the plasmid. The double digest with both HindIII and BamHI (lane 3) produces bands at 6kb and 1.2kb (red box), matching the backbone and insert, respectively. The results on the gel correspond to the predicted sizes.Restriction digest tips and tricks:
For your digest:
For your gel:

Source: https://blog.addgene.org/plasmids-101-how-to-verify-your-plasmid
0 Response to "How to Read Control/not-digested Plasmids After Electrophoretic Separation"
Post a Comment